Spectral Clustering
This is first in a series of Segmentation and Clustering articles. Clustering is a widely used unsupervised learning technique. ‘Spectral Clustering’ uses the connectivity approach to clustering.
Overview
Clustering is a widely used unsupervised learning technique. The grouping is such that the points in a cluster are similar to each other. ‘Spectral Clustering’ uses the connectivity approach to clustering, wherein communities of nodes (i.e. data points) that are connected or immediately next to each other are identified in a graph. The nodes are then mapped to a low-dimensional space that can be easily segregated to form clusters. Spectral Clustering uses information from the eigenvalues (spectrum) of special matrices (i.e. Affinity Matrix, Degree Matrix and Laplacian Matrix) derived from the graph or the data set.
Differences between Spectral Clustering and Conventional Clustering Techniques
Spectral clustering is flexible and allows us to cluster non-graphical data as well. It makes no assumptions about the form of the clusters. Clustering techniques, like K-Means, assume that the points assigned to a cluster are spherical about the cluster centre. This is a strong assumption and may not always be relevant. In such cases, Spectral Clustering helps create more accurate clusters. It can correctly cluster observations that actually belong to the same cluster, but are farther off than observations in other clusters, due to dimension reduction.
The data points in Spectral Clustering should be connected, but may not necessarily have convex boundaries, as opposed to the conventional clustering techniques, where clustering is based on the compactness of data points. Although, it is computationally expensive for large datasets, since eigenvalues and eigenvectors need to be computed and clustering is performed on these vectors. Also, for large datasets, the complexity increases and accuracy decreases significantly.
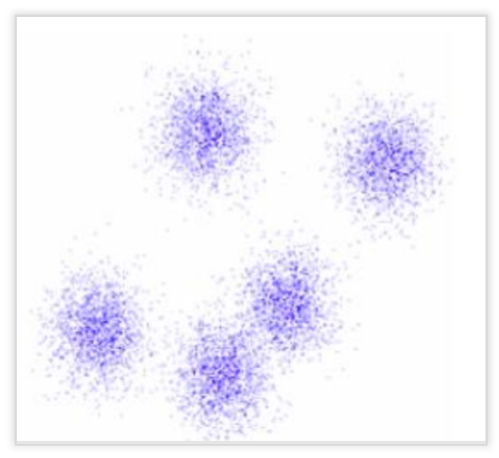
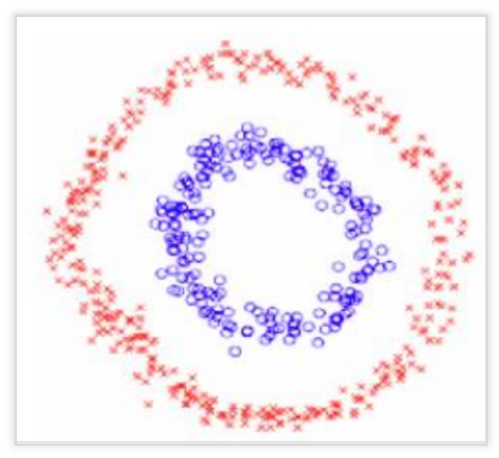
Basic Terminologies Used in Spectral Clustering
Adjacency and Affinity Matrix (A)
The graph (or set of data points) can be represented as an Adjacency Matrix, where the row and column indices represent the nodes, and the entries represent the absence or presence of an edge between the nodes (i.e. if the entry in row 0 and column 1 is 1, it would indicate that node 0 is connected to node 1). An Affinity Matrix is like an Adjacency Matrix, except the value for a pair of points expresses how similar those points are to each other. If pairs of points are very dissimilar then the affinity should be 0. If the points are identical, then the affinity might be 1. In this way, the affinity acts like the weights for the edges on our graph.
Degree Matrix (D)
A Degree Matrix is a diagonal matrix, where the degree of a node (i.e. values) of the diagonal is given by the number of edges connected to it. We can also obtain the degree of the nodes by taking the sum of each row in the adjacency matrix.
Laplacian Matrix (L)
This is another representation of the graph/data points, which attributes to the beautiful properties leveraged by Spectral Clustering. One such representation is obtained by subtracting the Adjacency Matrix from the Degree Matrix (i.e. L = D – A). To gain insights and perform clustering, the eigenvalues of L are used. Some useful ones are mentioned below:
- Spectral Gap: The first non-zero eigenvalue is called the Spectral Gap. The Spectral Gap gives us some notion of the density of the graph.
- Fiedler Value: The second eigenvalue is called the Fiedler Value, and the corresponding vector is the Fiedler vector. Each value in the Fiedler vector gives us information as to which side of the decision boundary a particular node belongs to.
- Using L, we find the first large gap between eigen values which generally indicates that the number of eigenvalues before this gap is equal to the number of clusters.
Cluster Eigenspace Problem’ Simplified
To find optimal clusters, the Laplacian matrix (L) should approximately be a block diagonal matrix, where each block represents a cluster. Each of these blocks consists of sub-blocks, that help us identify clusters with non-convex boundaries. The requirement of Spectral Clustering to find the actual cluster labels is such that the lowest eigenvalue and eigenvector pairs in the full space of the Laplacian Matrix should belong to different clusters. This happens when these eigenvectors correspond to the lowest eigenvectors in one of these sub-blocks present in the sub-spaces of the Laplacian Matrix. This restricts the eigenvalue spectrum of L, so that the set of lowest full-space eigenvalues consist of the lowest sub-block eigenvalues, which otherwise could have given us more meaningful insights of the graph/data points.
Emphasizing the Constraints on Real World Data
The constraint on the eigenvalue spectrum suggests that Spectral Clustering will only work on fairly uniform datasets; N uniformly sized clusters. Otherwise, there is no guarantee that the full space and sub-block eigen spectrums will line up nicely. In general, Spectral Clustering can be quite sensitive to changes in the similarity graph and to its parameters. Additionally, the selection of the similarity graph (i.e. K-Nearest Neighbours graph and e-neighbourhood graph) can affect results, as they respond differently to the data present in different densities and scales.
Furthermore, there can be consistency issues while using unnormalized Spectral Clustering algorithms, caused by the possible failure to converge the data points to suitable clusters as more data points are added. The A that is used to obtain the L is essential in determining similarity within data points, and hence, if A is not defined properly, poor similarity measures deter the applicability of Spectral Clustering. Unfortunately, there has been no systematic study which investigates the effects of the similarity graph and its parameters on clustering. Therefore, there are no well-justified rules of thumb.
Implementation in Python
Following is the link to the Jupyter Notebook which has the Spectral Clustering Implementation in Python:
References for further reading:
- Spectral Clustering: A Quick Overview:
https://calculatedcontent.com/2012/10/09/spectral-clustering/ - Spectral Clustering – Foundation and Application
https://towardsdatascience.com/spectral-clustering-aba2640c0d5b - Spectral Clustering from Scratch
https://medium.com/@tomernahshon/spectral-clustering-from-scratch-38c68968eae0 - A Tutorial on Spectral Clustering
https://arxiv.org/pdf/0711.0189.pdf
Technical articles are published from the Absolutdata Labs group, and hail from The Absolutdata Data Science Center of Excellence. These articles also appear in BrainWave, Absolutdata’s quarterly data science digest.
























